Humans would not have existed without viruses because viral protein plays a key role in the development of human embryo. However, at times, they pose existential threats in the form of diseases as in the case of current COVID-19 pandemic. Ironically, viruses comprise ~8% of our genome, that has been acquired during the course of evolution, making us “virtually a chimera”.
The most infamous and dreadful word of the year 2020 without doubt is ‘virus’. The novel coronavirus is responsible for the current unprecedented COVID-19 disease and an almost near collapse of the world economy. All this is caused by a tiny particle which is not even considered as ‘fully’ living because it is in a non-functional state outside the host, while only perpetuating inside upon infecting the host. More surprising and shocking is the fact that the humans have been carrying the viral “genes” since times immemorial and currently viral genes constitutes ~8% of the human genome (1). Just to put this in perspective, only ~1% human genome is functionally active responsible for making proteins that determine who we are.
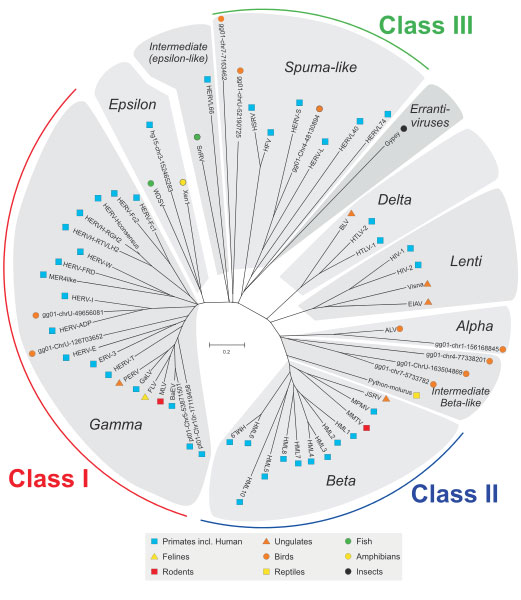
The story of relationship between humans and viruses started 20-100 million years ago when our ancestors got infected by viruses. Each endogenous retrovirus family is derived from a single infection of the germline cells by an exogenous retrovirus that after integrating into our ancestor, expanded and evolved (2). The propagation followed by the horizontal transfer from parents to offspring and today we have these viral genomes embedded in our DNA as human endogenous retroviruses (HERVs). This is a continuous process and may even be happening at the moment. Over the course of evolution, these HERVs acquired mutations, became stabilised in the human genome and lost their ability to cause the disease. The endogenous retroviruses are not only present in humans but are omnipresent in all living organisms. All these endogenous retroviruses grouped into three classes (Class I, II and III) occurring across different animal species exhibit a phylogenetic relationship based on their sequence similarity (3) as depicted in Figure below. HERVs belong to the Class I group.
Of the various embedded retroviruses present in the human genome, a classic example worth mentioning here, is that of a retroviral protein that is highly fusogenic envelope protein called syncytin, (5) whose original function in the virus was to fuse with host cells to cause infection. This protein has now been adapted in humans to form placenta (fusion of cells to make multinucleated cells) that not only provides food to foetus from the mother during pregnancy but also protects the foetus from the mother’s immune system due to the immunosuppressive nature of the syncytin protein. This particular HERV has proven to be beneficial to the human race by defining its very existence.
HERVs have also been implicated in providing innate immunity to the host by preventing further infection from related viruses or reducing the severity of the disease upon re-infection by similar type of viruses. A 2016 review by Katzourakis and Aswad (6) describes that endogenous viruses can act as regulatory elements for genes that control immune function, thereby leading to immunity development. In the same year, Chuong et al (7) demonstrated that certain HERVs act as regulatory enhancers by modulating the expression of IFN (interferon) inducible genes thereby providing innate immunity. HERV expression products can also act as pathogen-associated molecular patterns (PAMPs), triggering the cellular receptors responsible for host first line of defences (8-10).
Another interesting aspect of HERVs is that some of them show insertion polymorphisms, i.e. different number of copies are present in the genome due to insertional events. A study of 20 subjects belonging to different ethnic groups revealed insertion polymorphism patterns between 0-87% in all subjects (11). This can have implications in causing diseases by activation of certain genes that are otherwise silent.
Certain HERVs have also been shown to be associated with the development of autoimmune disorders such as multiple sclerosis (12). Under normal physiological conditions, HERV expression is tightly regulated while under pathological conditions due to changes in the external/internal environment, hormonal changes and/or microbial interaction can cause dysregulation of HERV expression, leading to disease.
The above characteristics of HERVs suggest that not only their presence in human genome is inevitable but they possess the ability to regulate the homeostasis of the immune system either by activating or suppressing it, thereby causing differential effects (from being beneficial to causing a disease) in hosts.
The COVID-19 pandemic is also caused by a retrovirus SARS-nCoV-2, that belongs to the influenza family, and it may be plausible that, during the course of evolution, genomes related to this family of viruses got integrated into the human genome and are now present as HERVs. It is surmised that these HERVs might exhibit different polymorphisms, as mentioned above, among people of different ethnicity. These polymorphisms may be in the form of differential copy number of these HERVs and/or presence or absence of mutations (changes in the genome sequence) accumulated over a period of time. This variability in the integrated HERVs may offer an explanation for the differential mortality rates and the severity of COVID-19 disease in different countries effected by the pandemic.
***
References:
1. Griffiths DJ 2001. Endogenous retroviruses in the human genome sequence. Genome Biol. (2001); 2(6) Reviews 1017. DOI: https://doi.org/10.1186/gb-2001-2-6-reviews1017
2. Boeke, J. D.; Stoye, J. P. (1997). “Retrotransposons, Endogenous Retroviruses, and the Evolution of Retroelements”. In Coffin, J. M.; Hughes, S. H.; Varmus, H. E. (eds.). Retroviruses. Cold Spring Harbor Laboratory Press. PMID 21433351.
3. Vargiu L, et al. Classification and characterization of human endogenous retroviruses; mosaic forms are common. Retrovirology (2016); 13: 7. DOI: 10.1186/s12977-015-0232-y
4. Classes_of_ERVs.jpg: Jern P, Sperber GO, Blomberg J (derivative work: Fgrammen (talk)), 2010. Available online at https://commons.wikimedia.org/wiki/File:Classes_of_ERVs.svg Accessed on 07 May 2020
5. Blond, JL; Lavillette, D; Cheynet, V; Bouton, O; Oriol, G; Chapel-Fernandes, S; Mandrandes, S; Mallet, F; Cosset, FL (7 April 2000). “An envelope glycoprotein of the human endogenous retrovirus HERV-W is expressed in the human placenta and fuses cells expressing the type D mammalian retrovirus receptor”. J. Virol. 74 (7): 3321–9. DOI: https://doi.org/10.1128/jvi.74.7.3321-3329.2000.
6. Katzourakis A, and Aswad A. Evolution: Endogenous Viruses Provide Shortcuts in Antiviral Immunity. Current Biology (2016). 26: R427-R429. http://dx.doi.org/10.1016/j.cub.2016.03.072
7. Chuong EB, Elde NC, and Feschotte C. Regulatory evolution of innate immunity through co-option of endogenous retroviruses. Science (2016) Vol. 351, Issue 6277, pp. 1083-1087. DOI: https://doi.org/10.1126/science.aad5497
8. Wolff F, Leisch M, Greil R, Risch A, Pleyer L. The double-edged sword of (re)expression of genes by hypomethylating agents: from viral mimicry to exploitation as priming agents for targeted immune checkpoint modulation. Cell Commun Signal (2017) 15:13. DOI: https://doi.org/10.1186/s12964-017-0168-z
9. Hurst TP, Magiorkinis G. Activation of the innate immune response by endogenous retroviruses. J Gen Virol. (2015) 96:1207–1218. DOI: https://doi.org/10.1099/vir.0.000017
10. Chiappinelli KB, Strissel PL, Desrichard A, Chan TA, Baylin SB, Correspondence S. Inhibiting DNA methylation causes an interferon response in cancer via dsRNA including endogenous retroviruses. Cell (2015) 162:974–986. DOI: https://doi.org/10.1016/j.cell.2015.07.011
11. Mehrab G, Sibel Y, Kaniye S, Sevgi M and Nermin G. Human endogenous retrovirus-H insertion screening. Molecular Medicine Reports (2013). DOI: https://doi.org/10.3892/mmr.2013.1295
12. Gröger V, and Cynis H. Human Endogenous Retroviruses and Their Putative Role in the Development of Autoimmune Disorders Such as Multiple Sclerosis. Front Microbiol. (2018); 9: 265. DOI: https://doi.org/10.3389/fmicb.2018.00265
***